Table of Contents |
The structure of DNA provided an intriguing hint about its mechanism of replication, as noted by James Watson and Francis Crick in their famous publication introducing their model (Watson and Crick, 1956). As you learned in the tutorial on DNA structure, nitrogenous bases bond according to complementary base pairing rules. This means that it is possible to determine the sequence of bases on one strand simply by knowing the sequence of bases on the other strand and provides a possible mechanism for replication of one DNA molecule to make two new molecules.
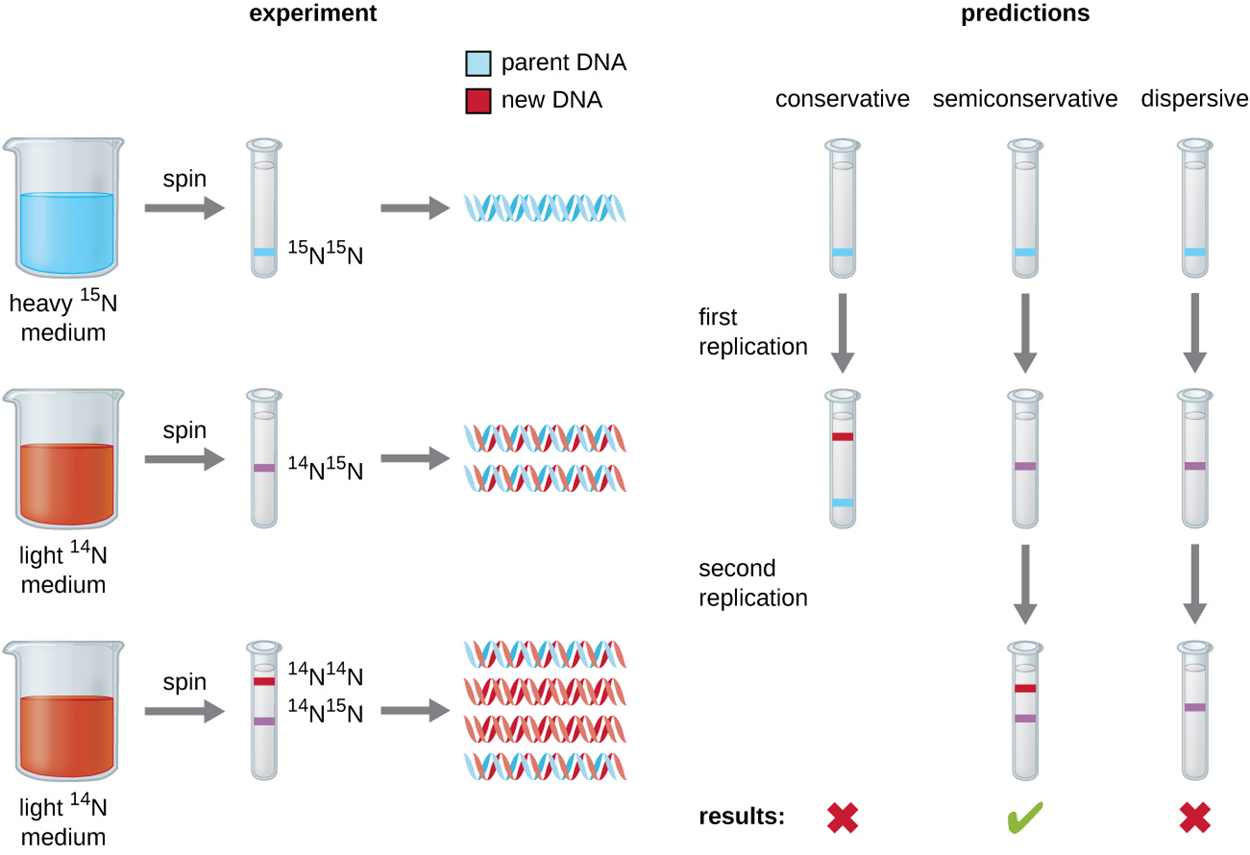
However, this simple overview betrays the complexity of the overall process. For example, the composition of daughter molecules was originally unclear. The image below shows three possible outcomes of replication. If replication were conservative, a parental double-stranded DNA (dsDNA) molecule would undergo replication to produce one daughter molecule identical to itself and one entirely new molecule. If replication were dispersive, then each daughter molecule would have two strands each composed of a mixture of parental and new nucleotides. If replication were semiconservative, then each parental molecule would produce two daughter molecules that each had one half of the parental molecule and one new half.
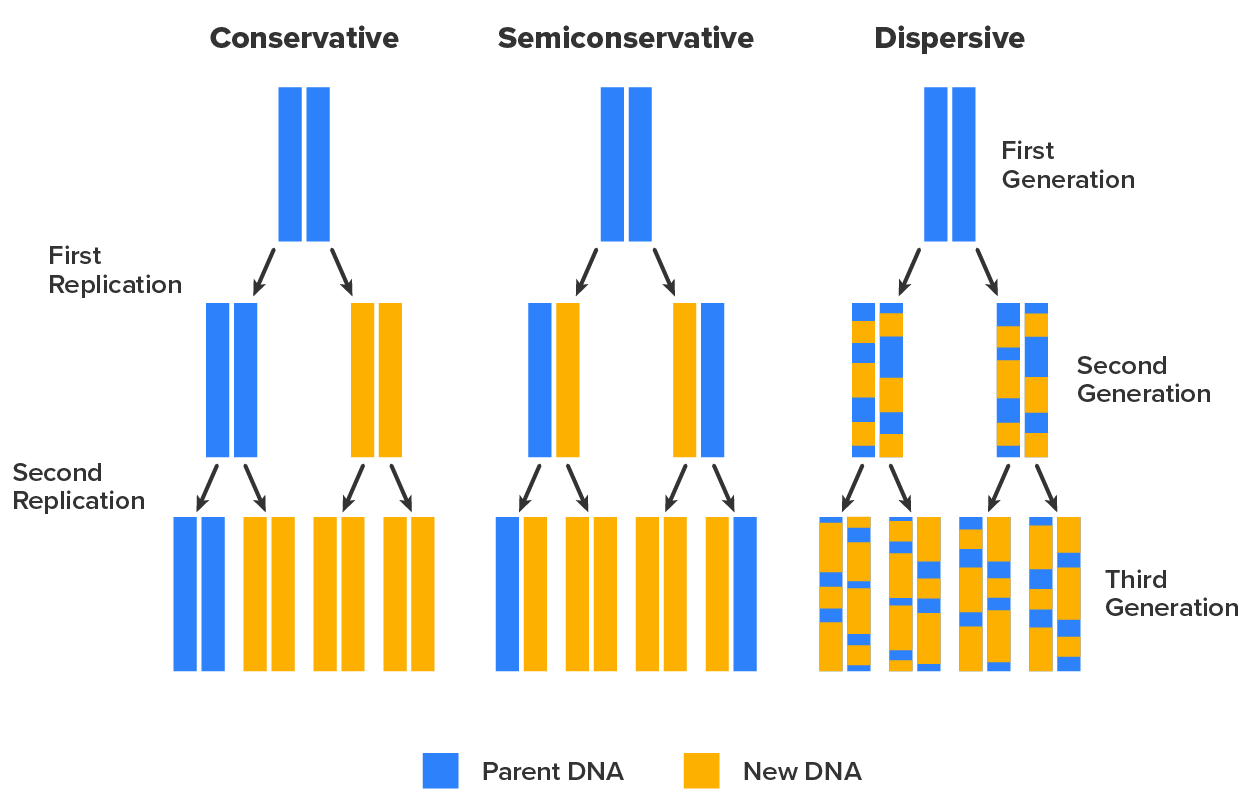
In 1958, Matthew Meselson (1930–Present) and Franklin Stahl (1929–Present) performed an experiment that determined that DNA replication was semiconservative. The experiment was designed so that there were three clearly distinct predictions based on whether DNA replication was conservative, semiconservative, or dispersive. The steps of the experiment were as follows.
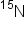 ) instead of the more common
) instead of the more common 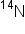 atoms.
atoms.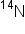 for a single generation, allowing the cells to produce daughter DNA molecules.
for a single generation, allowing the cells to produce daughter DNA molecules.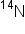 medium only and cells grown in
medium only and cells grown in 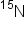 medium only.
medium only.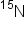 and DNA containing only
and DNA containing only 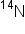 , indicating that it had one parental strand with
, indicating that it had one parental strand with 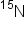 strand and one new strand containing
strand and one new strand containing 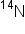 (i.e., it was semiconservative).
(i.e., it was semiconservative).
The image below shows the outcomes of centrifugation of the three types of samples as well as the predictions for each hypothesis: conservative, semiconservative, and dispersive. Only semiconservative replication produced results matching those obtained from the experiment after two replications of the DNA, producing an intermediate band (with one heavy band from the parent and one newly formed light band) in the first replication and then a mix of intermediate bands and light bands (from new DNA formed from the light strands of parents and new light strands) in the second replication. Conservative replication would have kept the original molecule with production of a new light molecule and dispersive replication would have mixed the strands so they would contain a mix of heavy and light nucleotides forming intermediate bands in the first and second replications.
Because bacteria have relatively small genomes, they replicate quickly and produce many mutants that can be utilized for research. This is why there has been considerable study of bacterial DNA replication. It only takes 42 minutes for an E. coli bacterium to replicate its entire single circular chromosome, which contains 4.6 million base pairs (Mbp).
Many enzymes and proteins are used in DNA replication, and these are summarized in a table at the end of the tutorial. These molecules prepare DNA for replication, help to hold components in place, and add components as needed.
One of the most important enzymes in this process is DNA polymerase. As its name suggests, DNA polymerase is an enzyme that produces polymers of DNA nucleotides. DNA-dependent DNA polymerases use DNA as a template, meaning that they add complementary nucleotides based on the DNA sequence (the unusual enzyme reverse transcriptase, which HIV uses to copy its RNA genome into cDNA, is an RNA-dependent DNA polymerase). There are several types of DNA polymerases. In prokaryotes, there are three types named using Roman numerals.
DNA polymerase III (also known as DNA pol III) is the enzyme that is responsible for the actual copying of one strand, whereas DNA pol I and DNA pol II are primarily required for repair. During replication, DNA pol III adds complementary base pairs to grow a new strand complementary to the template strand.
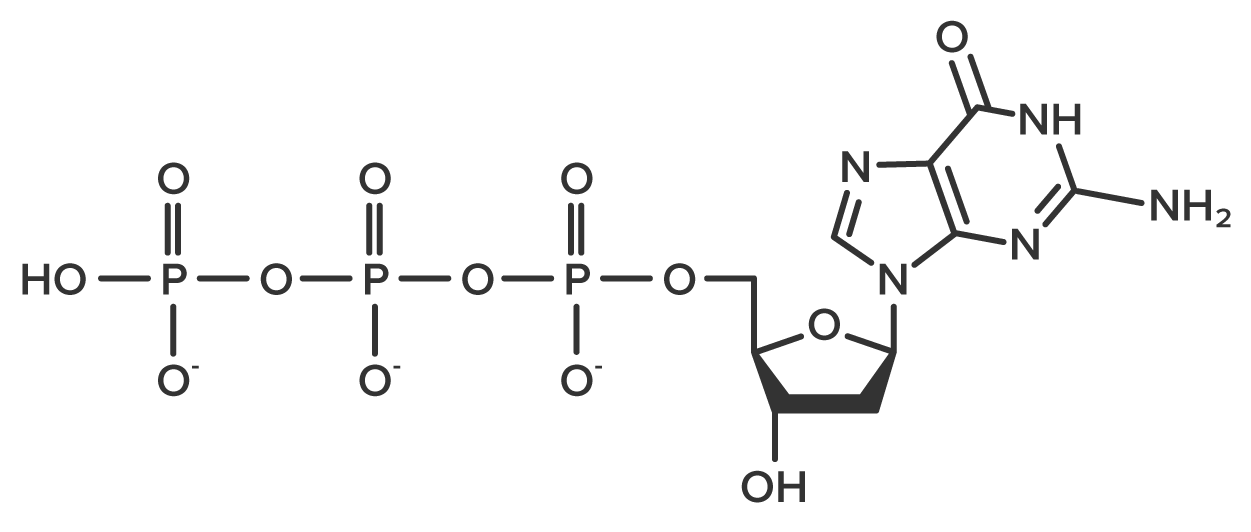
Each new nucleotide is added through the formation of a bond between phosphate and a free 3′-OH group. A triphosphate nucleotide, such as the guanosine triphosphate molecule shown below, undergoes bond cleavage to remove two phosphate groups. This exposes the phosphate that will form a new bond called the phosphodiester bond. Additionally, cleavage of the phosphate groups releases the energy needed for bond formation.
The process of replication is divided into three major steps: initiation, elongation, and termination. Initiation of replication occurs at a specific nucleotide sequence called the origin of replication, where various proteins bind to begin the replication process. Like most prokaryotes, E. coli has a single origin of replication.
For replication to occur, regions of DNA have to be made accessible. In chromosomes, DNA is packed around proteins to fit tightly within a cell. The process by which DNA is packed and unpacked through twisting is called supercoiling. It is important to prevent DNA from being too tightly or too loosely wound as other parts of the DNA are unwound for replication. Enzymes called topoisomerases act to make small nicks or cuts in DNA to resolve supercoiling, allowing replication to proceed. In many bacteria and some archaea, a specific topoisomerase called DNA gyrase is used.
Proteins are used to aid the organization of DNA. Specific proteins help bind DNA and aid in packaging. Other proteins bind to the origin of replication. In eukaryotes, DNA-binding proteins called histones are especially important in arranging DNA into compact chromatin. Prokaryotes do not have histones but do have similar proteins that are used to compact DNA in the nucleoid structure.
For bacterial DNA replication to begin, the supercoiled chromosome is relaxed by DNA gyrase. Another enzyme called helicase separates the DNA strands by breaking the hydrogen bonds between the nitrogenous base pairs. As the DNA opens up, it forms an open region called a replication bubble with Y-shaped replication forks at each end. The DNA near each replication fork is coated with single-stranded binding proteins to prevent the single-stranded DNA (ssDNA) from joining back together.
Once ssDNA is accessible at the origin of replication, nucleotides can be added to begin to form new strands. However, DNA pol III cannot start a new strand and can only add nucleotides in the 5′ to 3′ direction. Therefore, an enzyme called RNA primase adds a small sequence of RNA to start the new strand. RNA primase is an RNA polymerase, meaning that it builds polymers of RNA nucleotides.
Once the primer is in place, DNA pol III can begin to add DNA nucleotides and elongation can begin.
During elongation in DNA replication, the addition of nucleotides occurs at its maximal rate of about 1,000 nucleotides per second. The appearance of half of the replication fork during elongation is shown in the image below.
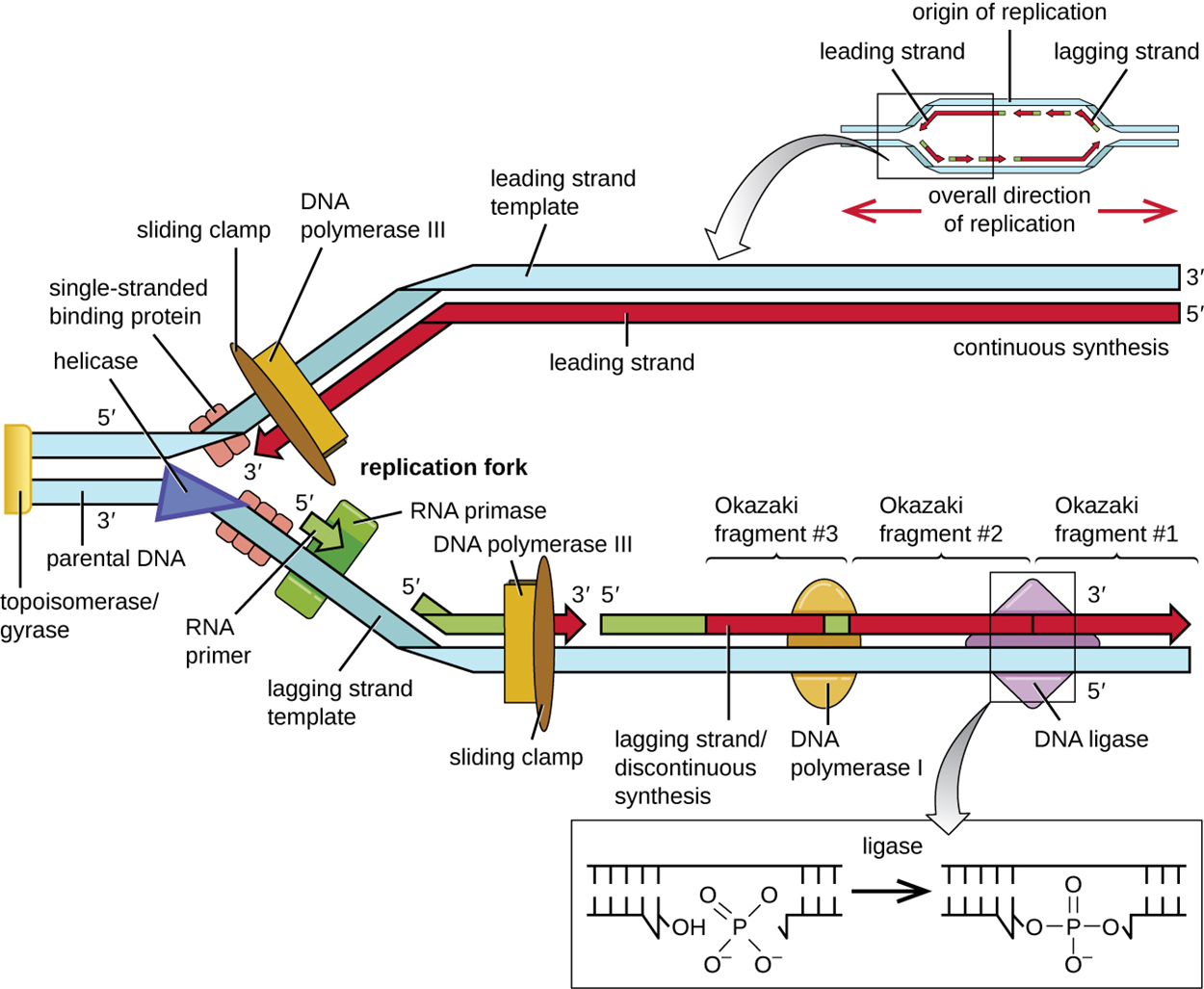
As you learned in the tutorial on DNA structure, a DNA double helix consists of antiparallel strands. Because DNA pol III can only add in the 5′ to 3′ direction, it means that it can only add continuously in one direction. The strands formed in this manner are called leading strands. To make new strands complementary to 5′ to 3′ parental DNA, new nucleotides are added moving away from (rather than toward) the replication fork. Making the other strands called lagging strands requires using small fragments of DNA called Okazaki fragments. Each Okazaki fragment requires its own primer. DNA pol III then adds nucleotides until it bumps into the existing strand and moves back again to begin a new fragment. Because of this process, lagging strands are said to be synthesized discontinuously.
Several things take place simultaneously. A protein called the sliding clamp holds the DNA polymerase in place as it continues to add nucleotides. Topoisomerase helps to prevent overwinding of the DNA double helix ahead of each replication fork by nicking the DNA backbone and resealing it. The RNA primers are removed by exonuclease activity of DNA pol I and then the gaps are filled in with DNA nucleotides. The gaps between the fragments are sealed by the enzyme DNA ligase, which catalyzes the formation of covalent phosphodiester linkages between the sides of each break.
Once the complete chromosome has been replicated, termination of DNA replication occurs. Less is known about this process than about initiation and elongation. Following replication, the resulting complete circular genomes of prokaryotes are concatenated, meaning that the circular DNA chromosomes are interlocked and must be separated from each other using cuts by bacterial topoisomerase IV. After the molecules have separated, topoisomerase reseals the cuts. This process is not necessary in eukaryotes, which have linear DNA molecules.
The table below summarizes the enzymes and proteins that work together during bacterial DNA replication.
| The Molecular Machinery Involved in Bacterial DNA Replication | |
|---|---|
| Enzyme or Factor | Function |
| DNA pol I | Exonuclease activity removes RNA primer and replaces it with newly synthesized DNA |
| DNA pol II | Primarily involved in repair; has 3′ to 5′ nuclease activity and may aid in repairing lesions |
| DNA pol III | Main enzyme that adds nucleotides in the 5′ to 3′ direction |
| Helicase | Opens the DNA helix by breaking hydrogen bonds between the nitrogenous bases |
| Ligase | Seals the gaps between the Okazaki fragments on the lagging strand to create one continuous DNA strand |
| Primase | Synthesizes RNA primers needed to start replication |
| Single-stranded binding proteins | Bind to ssDNA to prevent hydrogen bonding between DNA strands, reforming ds DNA |
| Sliding clamp | Helps hold DNA pol III in place when nucleotides are being added |
| Topoisomerase II (DNA gyrase) | Relaxes supercoiled chromosome to make DNA more accessible for the initiation of replication; helps relieve the stress on DNA when unwinding by causing breaks and then resealing the DNA |
| Topoisomerase IV | Introduces single-stranded break into concatenated chromosomes to release them from each other and then reseals the DNA |
DNA replication in eukaryotes has some important distinctions from DNA replication in prokaryotes. In part, this results from the fact that eukaryotes have much larger, more complex genomes arranged as multiple linear chromosomes. As shown in the image below, eukaryotes have multiple origins of replication on each chromosome that extend until they meet. Although this helps to reduce the time needed to replicate the entire genome, the rate of replication is approximately ten times slower than prokaryotic replication.
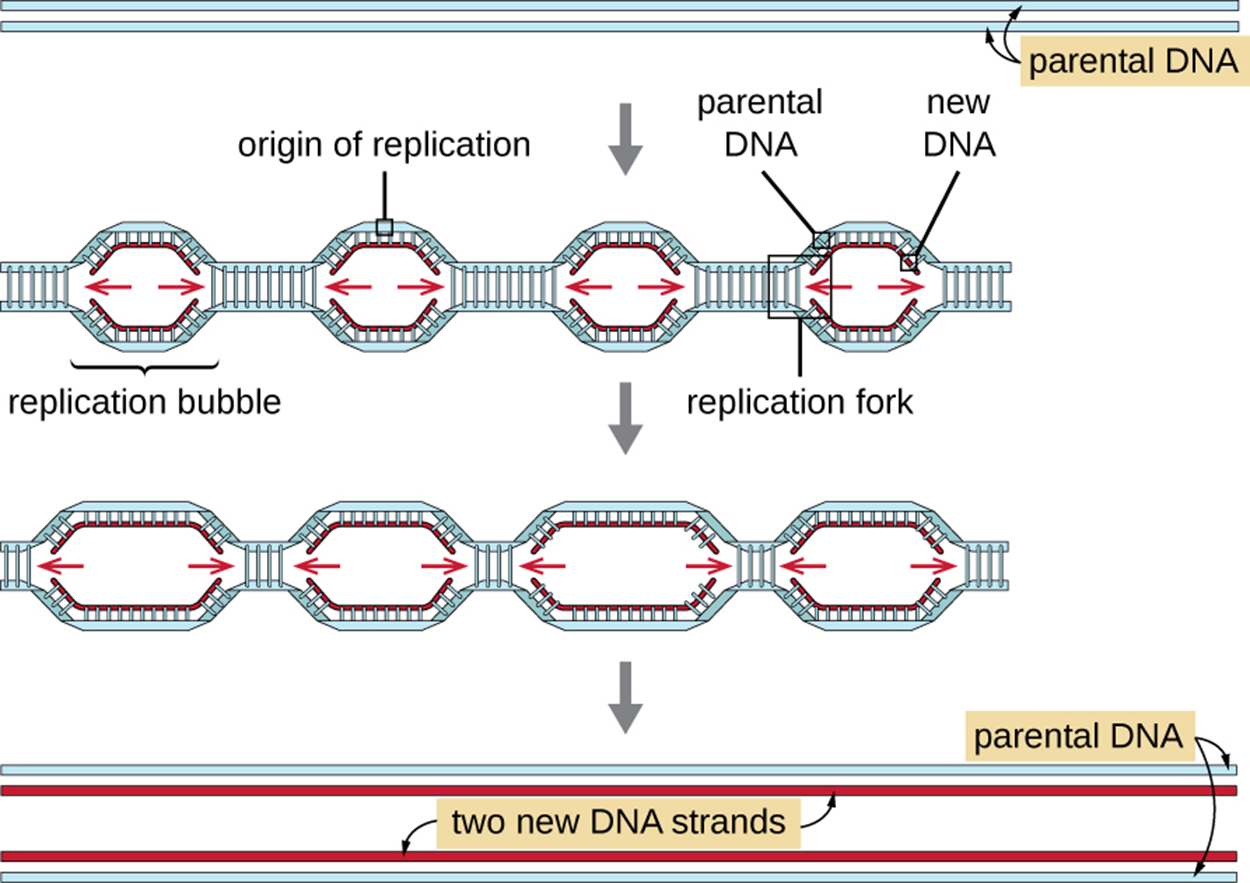
Despite these important differences, the most essential steps of replication are the same in prokaryotes and eukaryotes. Although there are more types of polymerases, the basic process of elongation using leading and lagging strands is the same.
A prereplication complex composed of several proteins, including helicase, forms at the origin of replication and recruits other components involved in the initiation of replication. These components include eukaryotic topoisomerase, single-stranded binding protein, RNA primase, and DNA polymerase.
Eukaryotic DNA polymerases are named using Greek letters. During elongation, DNA pol delta (δ) continuously synthesizes the leading strand and pol epsilon (ε) synthesizes the lagging strand. A sliding clamp protein holds DNA polymerase in place against the DNA. Instead of a DNA polymerase (used by bacteria), the enzyme ribonuclease H (RNase H) removes the RNA primer. After the primer is replaced with DNA nucleotides, DNA ligase seals the strands to stabilize them.
One important difference between prokaryotic and eukaryotic DNA replication results from the linear chromosomes of eukaryotes. As the image below shows, there is no way to make primers at the very end of linear chromosomes (noncoding repetitive sequences called telomeres). Over time, this means that the DNA becomes progressively shorter unless an enzyme called telomerase acts to fill the gaps as shown in the image.
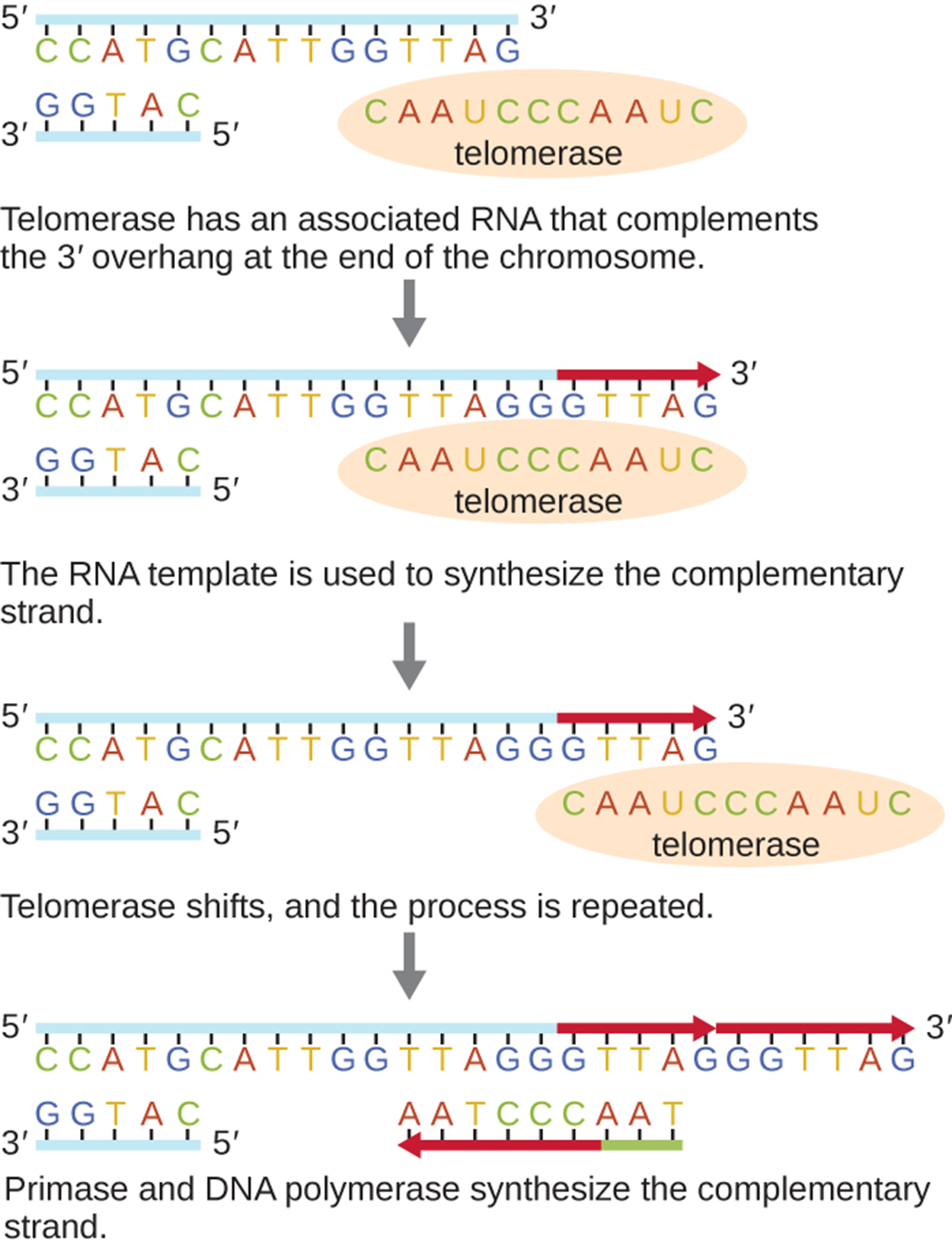
The table below summarizes some of the distinctions between bacterial and eukaryotic replication. Note that Archaea are not included in this table. Archaea have unique characteristics distinct from the other two domains but that is beyond the scope of this lesson (e.g., Sarmiento et al., 2014; Pérez-Arnaiz et al., 2020).
| Comparison of Bacterial and Eukaryotic Replication | ||
|---|---|---|
| Property | Bacteria | Eukaryotes |
| Genome structure | Single circular chromosome | Multiple linear chromosomes |
| Number of origins per chromosome | Single | Multiple |
| Rate of replication | 1,000 nucleotides per second | 100 nucleotides per second |
| Telomerase | Not present | Present |
| RNA primer removal | DNA pol I | RNase H |
| Strand elongation | DNA pol III | pol δ, pol ε |
A unique type of replication called rolling circle replication is used by some bacterial plasmids, some bacteriophages, and some eukaryotic viruses. This type of replication is only possible for DNA that is circular (as in plasmids) or capable of circularization (as in some viruses). The process in bacteria is summarized in the steps and image below.
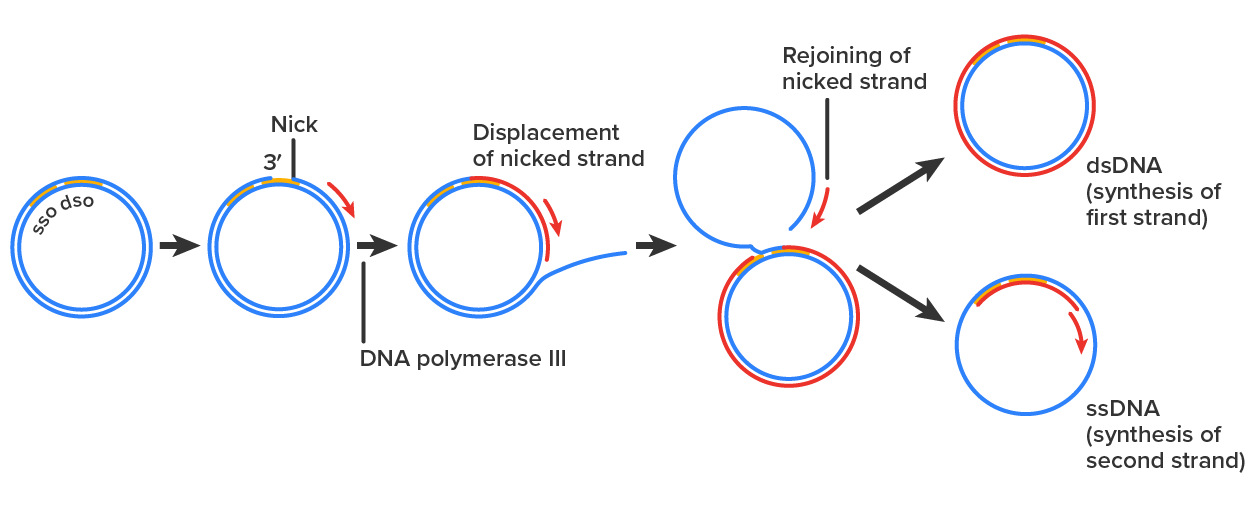
Source: THIS TUTORIAL HAS BEEN ADAPTED FROM OPENSTAX “MICROBIOLOGY.” ACCESS FOR FREE AT openstax.org/details/books/microbiology. LICENSE: CC ATTRIBUTION 4.0 INTERNATIONAL.
REFERENCES
Crick, F. H. & Watson, J. D. (1953). Molecular structure of nucleic acids: a structure for deoxyribose nucleic acid. Nature 171, 737–738. www.nature.com/articles/171737a0
Graham, M. K., & Meeker, A. (2017). Telomeres and telomerase in prostate cancer development and therapy. Nature reviews. Urology, 14(10), 607–619. doi.org/10.1038/nrurol.2017.104
Kuznetsova, A. A., Fedorova, O. S., & Kuznetsov, N. A. (2022). Structural and Molecular Kinetic Features of Activities of DNA Polymerases. International journal of molecular sciences, 23(12), 6373. doi.org/10.3390/ijms23126373
McKelvey, B. A., Umbricht, C. B., & Zeiger, M. A. (2020). Telomerase Reverse Transcriptase (TERT) Regulation in Thyroid Cancer: A Review. Frontiers in endocrinology, 11, 485. doi.org/10.3389/fendo.2020.00485
Pérez-Arnaiz, P., Dattani, A., Smith, V., & Allers, T. (2020). Haloferax volcanii-a model archaeon for studying DNA replication and repair. Open biology, 10(12), 200293. doi.org/10.1098/rsob.200293
Sarmiento, F., Long, F., Cann, I., & Whitman, W. B. (2014). Diversity of the DNA replication system in the Archaea domain. Archaea (Vancouver, B.C.), 2014, 675946. doi.org/10.1155/2014/675946